Research article
M.R. Al-Kaabi,1 W.U-Z. Tariq1 and A.A. Hassanein1
ازدياد مقاومة الجراثيم للمضادات الحيوية الشائعة في إمارة العين في دولة الإمارات العربية المتحدة
محمد راشد الكعبي، وحيد الزمان طارق، عواطف أبو حسنين
الخلاصـة: تَنْدُرُ المعلومات المحلِّية في إمارة العين في دولة الإمارات العربية المتحدة حول أنماط المقاومة للمضادات الحيوية. وفي هذه الدراسة الاستعادية أجرى الباحثون في أحد مستشفيات احالة الثالثية تحليلاً لنتائج حساسية المضادات الحيوية للأدوية على مدى خمس سنوات من عام 2004 حتى 2008، وقورِنَتْ مع دراسة سابقة أجريت في نفس المستشفى من عام 1999 حتى 2002. وتبين أن العنقودية الذهبية قد أظهرت انخفاضاً يُعتدّ به إحصائياً في حساسيتها للأوكساسيلين من 95.0% في الفترة 1999-2002 إلى %84.4 في عام 2008. وانخفضت حساسية الراكدة Acinetobacter للإيميبينيم من 99.0% في عام 2004 إلى 32.5% فقط في عام 2008. وظهرت في نفس الفترة مقاومة للسيفوكسيتين في حوالي نصف مُستفرَدات الإشريكية القولونية. كما اكتُشِفَ انخفاض يُعتدّ به إحصائياً في حساسية الزائفة الزنجارية بين عامي 1999 و2008 لجميع المضادات الحيوية التي جرى اختبارها تقريباً. ولم تُظهر الكلبسيلة أي تغير يُعتدّ به إحصائياً في المقاومة لأيٍّ من المضادات الحيوية التي جرى اختبارها. ويتبيَّن من الدراسة أن الحاجة ماسَّةٌ إلى بَذْل جهود حثيثة حتى يمكن تقليص اختطار انتشار ذراري الجراثيم المقاومة للمضادات الحيوية
ABSTRACT: There is a dearth of local information in Al Ain, United Arab Emirates about antibiotic resistance patterns. In this retrospective study in a tertiary referral hospital, antibiotic susceptibility results were analysed over the 5-year period 2004–08 and compared with a previous study in the same hospital during 1999–2002. Staphylococcus aureus showed a significant decrease in sensitivity to oxacillin from 95.0% in the period 1999–2002 to 84.4% in 2008. Sensitivity of Acinetobacter spp. to imipenem dropped from 99.0% in 2004 to only 32.5% in 2008. During the same period, almost half of Escherichia coli isolates developed resistance to cefotoxime. Significant reductions in sensitivity to Pseudomonas aeruginosa between 1999 and 2008 were found for almost all the antibiotics tested. Klebsiella spp. did not show any significant change in resistance to any of the tested antibiotics. Serious efforts are needed to reduce the risk of the spread of resistant strains of bacteria.
Augmentation de la résistance bactérienne aux antibiotiques courants à Al Ain, (Émirats arabes unis)
RÉSUMÉ: Il existe une pénurie d’informations locales à Al Ain (Émirats arabes unis) concernant les schémas de résistance aux antibiotiques. Dans la présente étude rétrospective conduite dans un hôpital de soins tertiaires, les résultats de sensibilité aux antibiotiques ont été analysés sur une période de 5 ans allant de 2004 à 2008 puis comparés aux résultats de l’étude précédente de 1999 à 2002 dans le même hôpital. La sensibilité de Staphylococcus aureus à l’oxacilline a diminué significativement, passant de 95,0 % entre 1999 et 2002 à 84,4 % en 2008. La sensibilité d’Acinetobacter spp. à l’imipénem a chuté, passant de 99,0 % en 2004 à seulement 32,5 % en 2008. Au cours de la même période, on a constaté un développement d’une résistance à la céfotoxime dans près de la moitié des isolats d’Escherichia coli. Des réductions importantes de la sensibilité de Pseudomonas aeruginosa entre 1999 et 2008 ont été observées pour presque tous les antibiotiques testés. La résistance de Klebsiella spp. aux antibiotiques testés est restée plutôt stable. Des efforts importants sont requis pour réduire le risque de propagation de souches bactériennes résistantes.
EMHJ, 2011, 17(6): 479-484
1Division of Microbiology, Unit of Laboratory Medicine, Tawam Hospital, Al Ain, United Arab Emirates (Correspondence to W.U-Z. Tariq:
Introduction
There is global concern about the growing problem of antimicrobial resistance, especially the appearance and rapid spread of methicillin-resistant Staphylococcus aureus (MRSA) in hospitals and the community [1]. The phenomenon is complex, involving specific microbial characteristics, selective pressures of antimicrobial use and demographic and technological changes that enhance the transmission of drug-resistant organisms. Such infections are becoming virtually untreatable and are leading to rising morbidity, mortality and health care costs [2].
In the United Arab Emirates (UAE), as in other countries, there is a growing concern about the indiscriminate prescription of antibiotics and even self-medication for minor ailments for which they many not be required. This can lead to the development of antibiotic-resistant strains of bacteria, thus reducing the effectiveness of certain antibiotics when they are required for a specific infection [3]. There has been growing concern in Saudi Arabia too as hospital strains of resistant bacteria are finding their way into the community: steps have been taken to modify local guidelines about the optimum use of antibiotics [4]. As there are variations in the resistance pattern in different countries, a thorough understanding of the changing local trends provides valuable information to hospitals to help them evaluate and plan for the proper management of clinical cases and to guide antibiotic stewardship programmes.
The present study at a tertiary referral hospital in Al Ain Emirate aimed to determine the susceptibility of certain bacterial isolates over the 5-year period 2004–08 and compare the results with a previous study in the same hospital carried out during the period 1999–2002. There is a dearth of local information in Al Ain and it was hoped that this study would provide valuable information about current patterns of resistance.
Methods
This retrospective study was carried out in Tawam hospital, Al Ain, over the period 2004–08. The susceptibility results of the following bacterial isolates were analysed: Staph. aureus, Escherichia coli, Pseudomonas aeruginosa, Klebsiella spp., Acinetobacter spp. and Stenotrophomonas maltophilia. The results were compared with a previous study carried out during the period 1999–2002 [5]. The previous study was performed in the same hospital, under similar conditions, but by different workers. In some cases, comparable data were not available for the same antibiotics/strains and the trends in susceptibility were traced during the present study only.
The bacteria were identified by conventional microbiology methods. Initially, between January 2004 and June 2006, the Vitek 1 microbiological culture analyser (BioMérieux) and the Analytic Profile Index strips (BioMérieux) were used for identifying bacteria. Subsequently, the Vitek 2 analyser replaced the Vitek 1. Antibiotic susceptibility was tested by the disk diffusion method during the period January 2004–June 2006, and then the Vitek 2 was used to perform antibiotic susceptibility testing for most isolates, while the disk diffusion method was retained for testing in case the Vitek 2 failed to produce satisfactory results or if retesting of susceptibility for a particular organism was considered necessary. The Vitek 2 provided minimum inhibitory concentration values for the tested isolates. The Clinical and Laboratory Standards Institute’s guidelines were used to interpret the antibiotic disk diffusion data [6].
Different antibiotic panels (as deemed appropriate, with the consensus of a microbiologist and the treating physicians) were applied for different organisms. The data were saved in a laboratory information system (Epicenter, version 4.0) which has the facility to eliminate patient duplicate specimens (i.e. repeat isolates from the same body site and hospital service) from analysis. The algorithm used for handling repeat isolates was patient-based and only the first isolate per patient was included in the analysis. The calculation of percentage susceptible did not include isolates with intermediate susceptibility. Cumulative antibiogram reports of the different isolates and antimicrobials for the period of 2004–08 were compared with the study in 1999–2002.
Logistic regression analysis was done using SPSS, version 17.0 to assess the association between year of sampling and the antimicrobial resistance of the tested pathogens to determine whether there were significant changes in the percentage resistance over time.
Results
Staph. aureus strains showed a steady decrease in sensitivity to oxacillin from 94.0% in 2002 to 84.4% in 2008. Comparing the average rate of sensitivity over the period 1999–2002 in the previous study (96.0%) with the rate in 2008 showed that the decrease was significant (P < 0.05) (Table 1).
Table 1 Antibiogram for isolates from a tertiary referral hospital in Al Ain by year
P. aeruginosa showed significant decreases in sensitivity (P < 0.05) between the average rate for 1999–2002 and the rate in 2008 to amikacin, cefepime, ciprofloxacin (88.9% to 81.6%), gentamicin, imipenem and piperacillin/tazobactam (Table 1). Sensitivity to azetreonam decreased significantly between 2004 and 2008 and to ceftazidime between 1999–2002 and 2007.
Acinetobacter spp. showed a significant decrease in sensitivity between 2004 and 2008 (P < 0.05) to all the tested antibiotics, imipenem, ceftazidime, ciprofloxacin, gentamicin and piperacillin/tazobactam (Table 1).
E. coli showed a significant decrease in sensitivity between the average for 1999–2002 study and 2008 (PE. coli did not show any significant change in resistance to nitrofurantoin, piperacillin/tazobactam, imipenem and trimethoprim/sulfamethoxazole over the period 2004–08.
Klebsiella spp. did not show any significant change in resistance to any of the tested antibiotics between 2004 and 2008 (Table 1).
There was a drop in sensitivity of Sten. maltophilia to sulfamethoxazole between 2004 and 08 but this was not significant (Table 1).
The number of extended spectrum of beta-lactabase (ESBL)-producing isolates found increased significantly over the period 2004–08 (P < 0.05) (Figure 1). The increase was evident for both isolates of E. coli (from 7.0% to 22.3%) and Klebsiella spp. (9.3% to 16.4%) (Figure 2).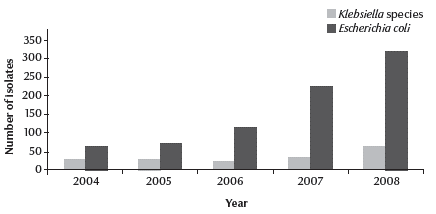
Figure 1 Total number of extended spectrum of beta-lactamase (ESBL)-producing isolates of Escherichia coli and Klebsiella spp. out of total ESBL isolates from a tertiary referral hospital in Al Ain for 2004–08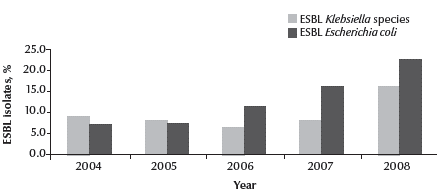
Figure 2 Percentage of extended spectrum of beta-lactamase (ESBL)-producing isolates of Escherichia coli and Klebsiella spp. from total ESBL isolates from a tertiary referral hospital in Al Ain for 2004–08
Discussion
Bacterial resistance to β-lactam antibiotics first appeared in the 1960s, as methicillin began to be used in clinical practice and was identified in 1% of isolates [7]. By 1993, serious concern was being expressed about the emerging problem that Japanese hospitals were experiencing due to MRSA. Nowadays, MRSA, with its serious implications, has appeared all over the world [8]. According to an estimate, the rate of MRSA strains have almost doubled in the United States of America (USA), during the same period, from 127 000 in 1999 to 278 000 in 2005. The number of people dying due to MRSA infections reached 17 000 in 2005, an increase from 11 000 in 1999 [9]. A similar trend had also been observed in the United Kingdom, where MRSA was reported to be a cause of death in 1652 cases in 2006, a rise from 51 MRSA-related deaths in 1993, albeit with a subsequent gradual decline in mortality [10]. The data for oxacillin in the present study showed that the rate of MRSA increased from 5.0% in the period 1999–2002 [5] to 15.6% in 2008 in the present study.
Acinetobacter spp. are also notorious for their ability to develop resistance to all classes of antimicrobials. A. baumannii is a hospital-acquired organism and is therefore exposed to a broad range of antibiotics. Most probably these genetic determinants have been acquired from other nonfermenter organisms in the environment, as well as common members of the Enterobacteriaceae. β-lactamases and efflux pumps have been implicated in the resistance mechanisms of Acinetobacter strains [11]. Over the years of our study, the sensitivity rates of Acinetobacter spp. to various antibiotics have than more halved and we have been left no good choice of antibiotics. Until recently, carbapenams would have been the choice against Acinetobacter spp. as our study showed that 99.0% of these organisms were sensitive to imipenem in 2004. By 2008, however, sensitivity had dropped to only 32.5%. Acinetobacter spp. commonly colonize patients in intensive care units, especially those who are intubated or have multiple intravenous lines or monitoring devices, postsurgical drains or urinary catheters. These organisms are found almost exclusively in patients admitted to hospitals. Irrespective of the fact that their role in disease causation is marginal and the cause of mortality and morbidity is due the underlying disease, their rising resistance against antimicrobial drugs is alarming [12].
E. coli is a ubiquitous organism, which may spread to human beings from animals, mainly poultry. It is becoming resistant to multiple drugs and the findings from poultry isolates are also reflected in the human isolates of this organism, mainly due to the extensive use of antibiotics in farming [12]. A study in the USA revealed that almost 331 of the 931 E. coli isolates from humans and poultry were resistant to 1 or more commonly used antibiotics [14]. During the same period, almost a quarter of E. coli isolates have developed resistance to cefotaxime. Moreover the organisms showing drug resistance have been shown to express high levels of virulence factors [15].
The problem of drug resistance in P. aeruginosa are multifold. The organism may accumulate intrinsic drug resistance mechanisms such as overproduction of cephalosporinase AmpC, increased drug efflux, fluoroquinolone target mutations and deficient production of porin OprD. Moreover, there are exogenous mechanisms such as production of secondary β-lactamases and aminoglycoside-modifying enzymes. These retain the ability to generate severe bloodstream infections. Therefore, multidrug-resistant P. aeruginosa may remain fully pathogenic [16]. We found significant reductions in sensitivity to P. aeruginosa between 1999 and 2008 for almost all the antibiotics tested.
Taking into consideration the local antibiogram data, guidelines have been issued for rational use of antibiotics in the UAE. The overuse of antibiotics is avoided and as far as possible the most appropriate antibiotic is used for an adequate period and not beyond. The overuse of antibiotic for surgical prophylaxis is also discouraged. Antibiotics are reserved for those patients who can benefit from them, i.e. those are the shown to be infected with the relevant bacterial pathogen. As far as possible, a narrow spectrum drug is used to cover a particular organism identified. Serious efforts are needed to reduce the risk of development of resistant strains of bacteria in the environment and in hospitals. Surveillance of resistance should be a key factor to generate data to inform improvements in clinical practice.
References
- Okuma K et al. Dissemination of new methicillin-resistant Staphylococcus aureus clones in the community. Journal of Clinical Microbiology, 2002, 40:4289–4294.
- Cohen ML. Epidemiology of drug resistance: implications for a post-antimicrobial era. Science, 1992, 257:1050–1055.
- Abasaeed A et al. Self-medication with antibiotics by the community of Abu Dhabi Emirate, United Arab Emirates. Journal of Infection in Developing Countries, 2009, 3(7):491–497.
- Shibl A. The problem of antibiotic resistance. Arab Health Magazine, 2007, 12:20–21.
- Jumaa PA, Neringer R. A survey of antimicrobial resistance in a tertiary referral hospital in the United Arab Emirates. Journal of Chemotherapy (Florence, Italy), 2005, 17:376–379.
- Performance standards for antimicrobial susceptibility testing: nineteenth informational supplement M100-S19. Wayne, Pennsylvania, Clinical and Laboratory Standards Institute, 2009.
- Parker MT, Hewitt JH. Methicillin resistance in Staphylococcus aureus. Lancet, 1970, 295:800–804.
- Donovan J. Antibiotic resistance in Australia. Health Issues No. 69, December 2001:1–5.
- Klein E, Smith DL, Laxminarayan R. Hospitalizations and deaths caused by methicillin-resistant Staphylococcus aureus, United States, 1999–2005. Emerging Infectious Diseases, 2007, 13:1840–1846.
- Jarvis WR. Prevention and control of methicillin-resistant Staphylococcus aureus: dealing with reality, resistance, and resistance to reality. Clinical Infectious Diseases, 2010, 50(2):218–220.
- Gootz TD, Marra A. Acinetobacter baumannii: an emerging multidrug-resistant threat. Expert Review of Anti-Infective Therapy, 2008, 6:309–325.
- Silvia Munoz-Price LS, Weinstein RA. Acinetobacter infection. New England Journal of Medicine, 2008, 358:1271–1281.
- Johnson JR et al. Similarity between human and chicken Escherichia coli isolates in relation to ciprofloxacin resistance status. Journal of Infectious Diseases, 2006, 194:71–78.
- Johnson JR et al. Antimicrobial drug-resistant Escherichia coli from humans and poultry products, Minnesota and Wisconsin, 2002–2004. Emerging Infectious Diseases, 2007, 13:838–846.
- Sharma S, Bhat GK, Shenoy S. Virulence factors and drug resistance in Escherichia coli isolated from extraintestinal infections. Indian Journal of Medical Microbiology, 2007, 25:369–373.
- Hocquet D et al. Pseudomonas aeruginosa may accumulate drug resistance mechanisms without losing its ability to cause bloodstream infections. Antimicrobial Agents and Chemo–therapy, 2007, 51:3531–3536.








